Software
The core facility uses a number of different applications for the data analysis, including Affymetrix software and Illumina software. We also use freeware like R / Bioconductor and various online resources. The commercial applications we are currently using are listed below.
| Partek Genomic Suite Partek GS is a comprehensive suite of advanced statistics and interactive data visualization. It is supporting all microarray and next generation sequencing technologies including DGE and gene expression, RNA-seq and alternative splicing, copy number and association, ChIP-chip, ChIP-seq, and microRNA. |
 |
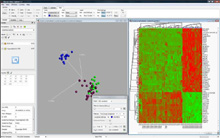
| Qlucore Omics Explorer Provides a tool to visualize and explore large microarray and real-time PCR data sets such as gene expression, microRNA, DNA methylation, protein expression and any multivariate data of size up to 1000 x 100,000. |
 |
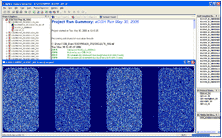
| Agilent Feature Extraction Agilent's Feature Extraction software reads and processes up to 100 raw microarray image files in an automated, walkaway mode. The software automatically finds and places microarray grids, rejects outlier pixels, accurately determines feature intensities and ratios, flags outlier pixels, and calculates statistical confidences. |
 |
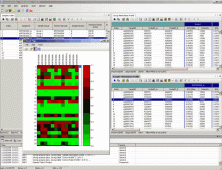
| Illumina GenomeStudio The GenomeStudio Software package comprises seven discrete application modules that enable you to conveniently compare data from different applications to obtain a comprehensive view of the genome, gene expression, and regulation. The software's modular nature allows new applications to be added as necessary. Data generated by Illumina's platforms can be easily exported to a number of third-party software tools for more secondary or tertiary analysis through the use of plug-ins integrated within the GenomeStudio module architecture. |
 |

- Customer Information:
- Project Submission
- External Users
- iLabs
- NGS Sample Submission
- How to acknowledge BEA
- Reference List
- Latest Newsletter
- FAQs
- BEA Price List:
- Order Forms:
- Links:
- KI core facilities for research
- SICOF-Single cell core facility
- CBB-Centre for Bioinformatics and Biostatistics
- SciLifeLab
- National Bioinformatics Institute Sweden
- Resources:
| BEA Service Fees |  |
| BEA Services |  |  |

